RESEARCH PROJECTS
Brain Computer Interfaces
Brain computer interface (BCI) technology is an emerging human computer interaction modality that involves invasive or noninvasive brain activity sensing coupled with signal processing and intent classification for the purpose of providing an interface to the operator that will enable communication with and via computers, control of external devices, computers, and robotic agents including virtual avatars. The technology can benefit both persons with various levels of physical impairments and the general population in certain applications. Nearly two million Americans have severe motor and speech impairments that require the use of Augmentative and Alternative Communication (AAC) devices. Communication rates achieved through AAC devices range from 6-15 words per minute for touch-based methods and 2-8 letters per minute for state-of-the-art BCI. There is huge gap in communication efficiency between the typical spoken communication and the ones achieved by the AAC devices. Our BCI research is supported by NSF, NIH, DARPA, and NLMFF.
RSVP Keyboard
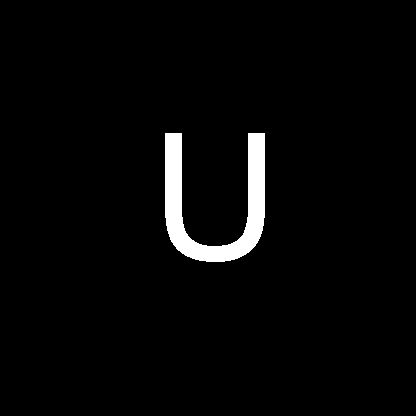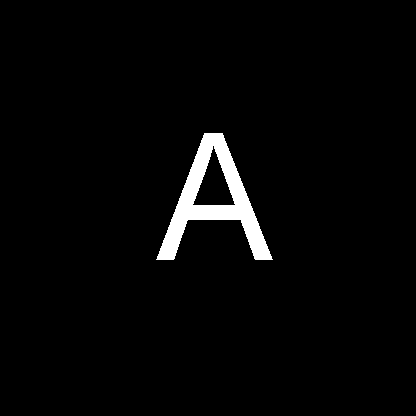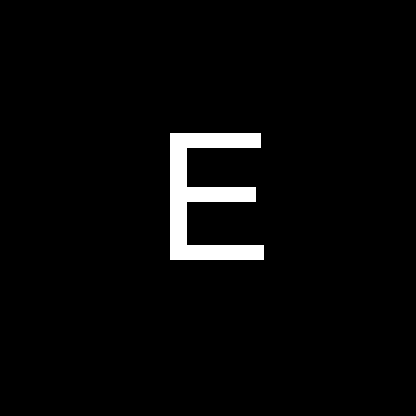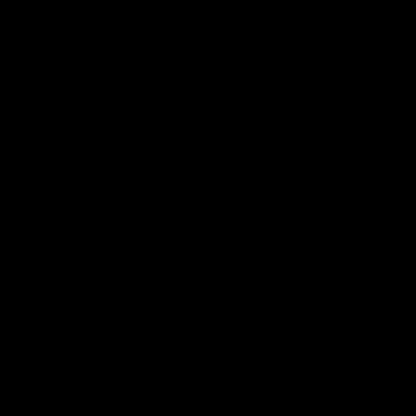
People with severe speech and physical impairments can benefit from a direct brain computer interface for their communication needs. This project aims to develop an AAC interface using noninvasive EEG sensors to infer the user's intent regarding desired letters and symbols during text generation. The designed RSVP Keyboard system will utilize rapid serial visual presentation of letter sequences coupled with probabilistic and adaptive open vocabulary language models and EEG signal processing and classification algorithms. The designed brain interface relies on event related potentials including the P300 signal. The project design tightly couples feedback from locked-in consultants who will test the system design at regular intervals and provide critical feedback in future design improvements.
RSVP iconCHAT
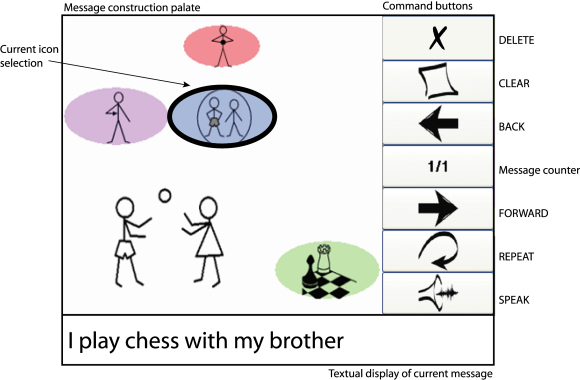
Most existing BCI research for assistive communication interfaces focus on the use letters to generate written text. This requires the subjects to be literate and comfortable with typing in general. As an alternative approach, in this project, we seek to design a BCI that enables the user to generate language using an iconic language called iconCHAT developed by Dr. Rupal Patel. Expected benefits include increased speed of communication as each icon might represent a word or phrase so equal number of item selection can construct a sentence instead of a word compared to RSVP Keyboard. A potential drawback is the necessity for the subject to become familiar with the icon set; this also means a closed vocabulary language generation system which is suitable for limited context interactions. An option to switch to letters for open vocabulary text generation can be incorporated to the final design for users that prefer to have this option.
Augmented Cognition Computer Games
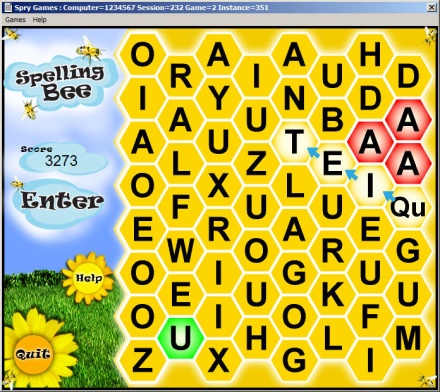 This
project aims to develop interactive computer games that are aware of the
user's cognitive state and engagement in order to adjust its challenge
level automatically for the purpose of maintaining a predetermined
effective mental effort level. The system features a fusion of passive
BCI that assesses the neurological markers in relationship to behavioral
markers as well as performance indices specifically designed to measure
cognitive engagement. These customized games are specifically designed
to be simultaneously entertaining and engaging in a manner that emulates
established batteries of cognitive tests and exercises.
This
project aims to develop interactive computer games that are aware of the
user's cognitive state and engagement in order to adjust its challenge
level automatically for the purpose of maintaining a predetermined
effective mental effort level. The system features a fusion of passive
BCI that assesses the neurological markers in relationship to behavioral
markers as well as performance indices specifically designed to measure
cognitive engagement. These customized games are specifically designed
to be simultaneously entertaining and engaging in a manner that emulates
established batteries of cognitive tests and exercises.
SSVEP-based BCI Design

Steady state visually evoked potentials (SSVEP) of the visual cortex provide an alternative means for enabling communication between a brain and a computer system. Being relatively easy to induce, and requiring almost no subject training and minimal calibration time, SSVEP signals are prime candidates for developing BCI controlled applications. We are exploring the use of SSVEP-inducing stimuli and associated signal processing and classification algorithms primarily in environmental control applications such as navigating a robot/wheelchair. The reliable and robust nature of these signals also make them ideal for undergraduate research activities in BCI technology and application development .
Biomedical Image Processing
Biomedical image processing applications present great challenges for statistical inference, machine learning, and pattern recognition. Our group focuses on a variety of contemporary problems that emerge in radiation therapy, basic neuroscience, and biology. These efforts are supported by NSF, Massachusetts General Hospital, and start-up funds provided by Northeastern University.
Real Time Lung Tumor Tracking
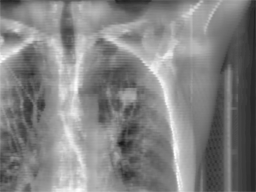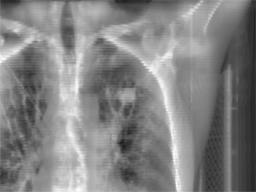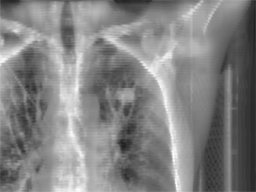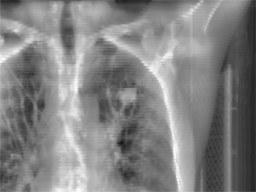 Radiotherapy
is an effective treatment technique for lung cancer. However, the movement
of lung tumors during normal respiration makes it difficult to accurately
irradiate the tumor. Precise lung tumor localization is vital to
efficiently treating the tumor and avoiding unnecessary radiation exposure
of normal tissues. Estimating the motion model of the tumor may lead to
improved treatment planning and dose calculation throughout the therapy.
4D Computed Tomography (CT) images taken prior to treatment (to develop
the patient's treatment plan) provide valuable information about the
movement of the thoracic organs and the tumor. For each radiotherapy
treatment session, a set of kilovoltage X-ray images (kV images) are
acquired to aid in the alignment of the treatment target relative to the
radiation beam. This project focuses on
real time lung tumor tracking by incorporating information and labels from
4DCT on kV x-ray videos for treatment-day specific tumor motion models.
Radiotherapy
is an effective treatment technique for lung cancer. However, the movement
of lung tumors during normal respiration makes it difficult to accurately
irradiate the tumor. Precise lung tumor localization is vital to
efficiently treating the tumor and avoiding unnecessary radiation exposure
of normal tissues. Estimating the motion model of the tumor may lead to
improved treatment planning and dose calculation throughout the therapy.
4D Computed Tomography (CT) images taken prior to treatment (to develop
the patient's treatment plan) provide valuable information about the
movement of the thoracic organs and the tumor. For each radiotherapy
treatment session, a set of kilovoltage X-ray images (kV images) are
acquired to aid in the alignment of the treatment target relative to the
radiation beam. This project focuses on
real time lung tumor tracking by incorporating information and labels from
4DCT on kV x-ray videos for treatment-day specific tumor motion models.
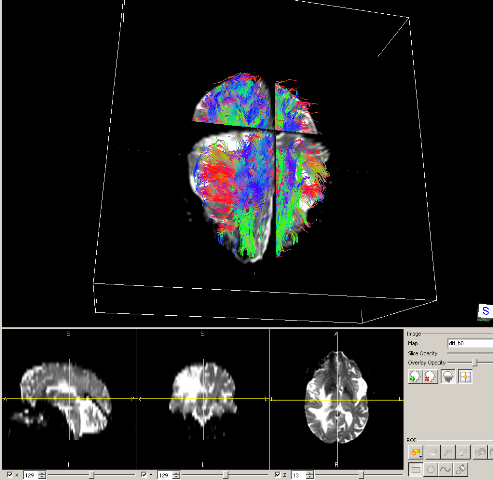
Predicting Migration of Cancer Cells using Diffusion Tensor Imaging
Diffusion tensor Magnetic Resonance Imaging (MRI) is a biomedical visualization technique, which shows direction of water diffusion. Diffusion Tensor Imaging (DTI) is an application of diffusion MRI aiming to extract and characterize diffusion anisotropy affects. When applied to brain, DTI shows the motion flow in the white matter of brain. Radiotherapy is widely used intreatment of malignant brain lesions. Current treatment plans include a fixed length margin around the tumor region of interest to account for the possible tumor spread. However, progressive tumors might occur outside of these margins, which requires more analysis about the spread of brain tumors. In this project, we are interested in predicting migration of cancer cells by using diffusion tensor MRI and finding out whether neuronal fiber pathways might provide possible routes for the spread of cancer cells. The outcome will help creating improved anisotropic margins for radiation treatment of brain tumors.
Tractography Techniques for Tubular Structures with Automated Bifurcation Detection
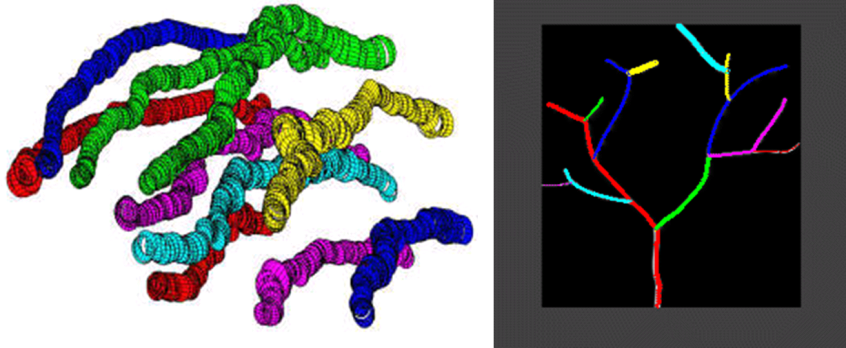
Tree and graph structures consisting of tubular segments are omnipresent in biological systems; consider the axons and dendritic trees of neurons, vascular networks in various organs, bronchial tree in the lungs, river and road networks in geospatial imagery (a 2-dimensional instantiation of the same concept). Biomedical 3-dimensional (3D) imagery obtained using various imaging modalities including confocal microscopy, computerized tomography (CT), magnetic resonance imaging (MRI) can be utilized to analyze and extract the 3D structure of tubular object networks which could be used to model and simulate such networks structurally and functionally, as well as design virtual endoscopy type diagnostic applications. We are currently focusing on confocal microscopy imagery of neural networks and their dendritic tree structures. Especially, we have been working with Brainbow imagery. Automatic extraction of dendritic tree structures is an important first step in processing massive image data to be used in improving our understanding of how the brain and the nervous system works.
Semi-supervised Segmentation of Organs and Tumors in 4D-CT
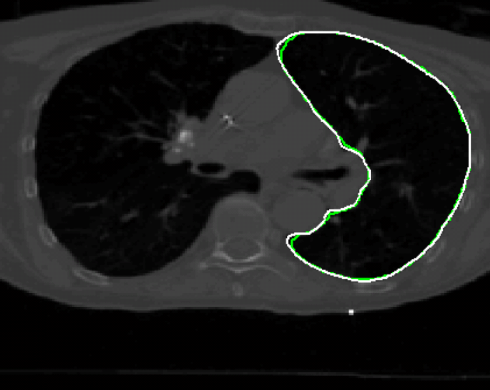 Radiation
therapy of tumors relies on 3D-CT and 4D-CT treatment planning imagery
which is manually segmented (delineated) and labeled prior to the
optimization of the dose delivery plan. Automatic segmentation of all
regions of interest has been a topic of intense research interest yet
clinical practitioners are unsatisfied with the performance of existing
solutions. It is our goal to keep the clinician in the loop during the
process and design an interactive segmentation tool that fuses inputs
from the expert user with low level features in imagery to speed up the
manual delineation process. One important goal is to accurately transfer
object boundaries from one respiration phase of 4D-CT to the other
without performing 3D-to-3D deformable registration.
Radiation
therapy of tumors relies on 3D-CT and 4D-CT treatment planning imagery
which is manually segmented (delineated) and labeled prior to the
optimization of the dose delivery plan. Automatic segmentation of all
regions of interest has been a topic of intense research interest yet
clinical practitioners are unsatisfied with the performance of existing
solutions. It is our goal to keep the clinician in the loop during the
process and design an interactive segmentation tool that fuses inputs
from the expert user with low level features in imagery to speed up the
manual delineation process. One important goal is to accurately transfer
object boundaries from one respiration phase of 4D-CT to the other
without performing 3D-to-3D deformable registration.
Interaction of Nanoparticles with Lipid Vesicles
 Novel
properties of nanoparticles have numerous potential technological
applications but at the same time underlie new kinds of biological
effects. Uniqueness of nanoparticles and nanomaterials requires a new
experimental methodology in order to acquire knowledge about their toxic
effect on biological systems and environment in general. Much evidence
suggests that nanoparticles affect cell membrane stability and
subsequently exert toxic effects. To explore these effects, we propose
an in vitro experiment with a simple biological system, a simulation of
cell membrane - lipid vesicle. Giant lipid vesicles are a promising
choice due to controllability and repeatability of experimental
conditions. Due to their size, which is on the same order of magnitude
as the size of cells, vesicles can be directly observed under the light
microscope. Vesicles' shape changes and fluctuations have been widely
investigated by various techniques, however microscopy imaging is among
most common. Most researchers focus on recording single vesicles
throughout the exposure period and observing changes in their
morphology. Instead, we propose recording populations of vesicles at
various times of exposure and calculating statistical analysis of the
vesicles size and morphological distributions. The hypothesis is that
different nanoparticles have different potential to interfere with lipid
membranes. We expect that this will be manifested as vesicles' shape
transformation. Our preliminary experiments already showed that changes
in populations of vesicles can be detected with this approach. Our
future direction is toward improving existing approaches to capturing of
images and their post processing.
Novel
properties of nanoparticles have numerous potential technological
applications but at the same time underlie new kinds of biological
effects. Uniqueness of nanoparticles and nanomaterials requires a new
experimental methodology in order to acquire knowledge about their toxic
effect on biological systems and environment in general. Much evidence
suggests that nanoparticles affect cell membrane stability and
subsequently exert toxic effects. To explore these effects, we propose
an in vitro experiment with a simple biological system, a simulation of
cell membrane - lipid vesicle. Giant lipid vesicles are a promising
choice due to controllability and repeatability of experimental
conditions. Due to their size, which is on the same order of magnitude
as the size of cells, vesicles can be directly observed under the light
microscope. Vesicles' shape changes and fluctuations have been widely
investigated by various techniques, however microscopy imaging is among
most common. Most researchers focus on recording single vesicles
throughout the exposure period and observing changes in their
morphology. Instead, we propose recording populations of vesicles at
various times of exposure and calculating statistical analysis of the
vesicles size and morphological distributions. The hypothesis is that
different nanoparticles have different potential to interfere with lipid
membranes. We expect that this will be manifested as vesicles' shape
transformation. Our preliminary experiments already showed that changes
in populations of vesicles can be detected with this approach. Our
future direction is toward improving existing approaches to capturing of
images and their post processing.
TOPICS OF THEORETICAL INTEREST
Tensor Decomposition Analysis
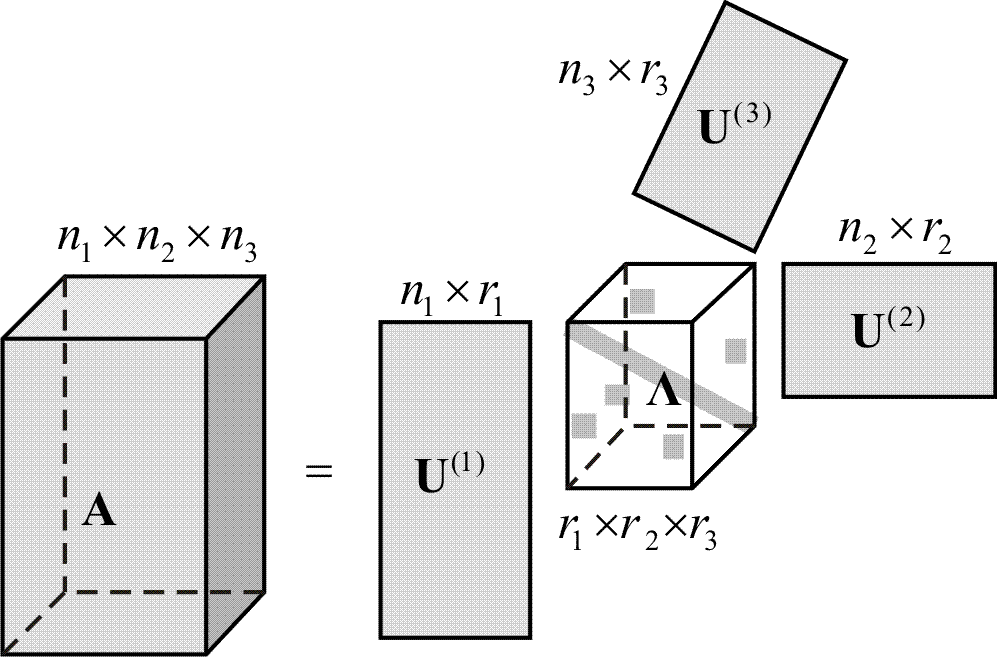 A
tensor is a multiway array of scalars; in this context it is a
generalization of scalars (order-0), vectors (order-1), and matrices
(order-2) to higher-order structures. Tensor analysis is an increasingly
relevant and interesting field of inquiry in signal processing and
machine learning as generalizations of rank-1 decompositions of matrices
such as the singular value decomposition (or eigendecomposition of
symmetric matrices) have found considerable application and success.
Decompositions of tensors emerge and become relevant when high-order
data or statistics are analyzed. Canonical decomposition (also called
parallel factor analysis) and other decomposition methodologies exist
and exhibit useful properties such as uniqueness. However, this
particular decomposition lacks structure in its vector frame that forms
the rank-one decomposition components, which prevents recursive solution
formulations as in deflation of principal components. The goal of this
project is to develop a non-redundant tensor decomposition, which can be
interpreted as a generalization of the spherical coordinate system of
vectors and the orthogonal matrix group based on a predetermined frame
of basis vectors. This imposed structure prevents the decomposition from
achieving the minimum cross-product tensor rank as canonical
decomposition does; however, we think it provides a solution that is
more suitable for recursive calculations that is useful and important in
dimension reduction applications.
A
tensor is a multiway array of scalars; in this context it is a
generalization of scalars (order-0), vectors (order-1), and matrices
(order-2) to higher-order structures. Tensor analysis is an increasingly
relevant and interesting field of inquiry in signal processing and
machine learning as generalizations of rank-1 decompositions of matrices
such as the singular value decomposition (or eigendecomposition of
symmetric matrices) have found considerable application and success.
Decompositions of tensors emerge and become relevant when high-order
data or statistics are analyzed. Canonical decomposition (also called
parallel factor analysis) and other decomposition methodologies exist
and exhibit useful properties such as uniqueness. However, this
particular decomposition lacks structure in its vector frame that forms
the rank-one decomposition components, which prevents recursive solution
formulations as in deflation of principal components. The goal of this
project is to develop a non-redundant tensor decomposition, which can be
interpreted as a generalization of the spherical coordinate system of
vectors and the orthogonal matrix group based on a predetermined frame
of basis vectors. This imposed structure prevents the decomposition from
achieving the minimum cross-product tensor rank as canonical
decomposition does; however, we think it provides a solution that is
more suitable for recursive calculations that is useful and important in
dimension reduction applications.
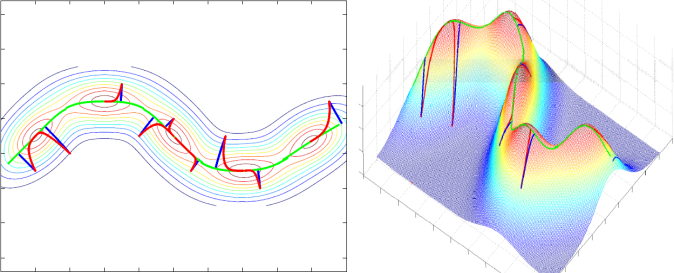
Principal Curves and Surfaces
Principal, saddle, and minor surfaces are critical surfaces of various dimensions that underlie functions which represent data density over some suitable space; this could be color or other feature distribution over space for images or probability density for arbitrary features in a statistical machine learning setting. We present a generalization of the local maximum, minimum, or saddle point concept as the fundamental definition of such surfaces in an attempt to frame the problem of manifold learning (and related problems with various names) as a geometrical structure identification problem. Our definition of the principal surface of a function is as follows: a point is an element of the principal surface if that point is a local maximum of the function in the orthogonal subspace at that point. This modifies Hastie's definition of self-consistent principal surfaces by replacing the mean-in-the-orthogonal-subspace with local maximum. The advantages are: (i) the definition is local and is only concerned with local derivatives of the function, (ii) it is trivial to define other critical surfaces such as saddle and minor surfaces, which generalize critical points (0-dimensional critical surfaces) to any dimensionality, (iii) since given the function (for instance the pdf) we can calculate the gradient, Hessian, etc at any point, we can know whether a point is on this principal surface or not without solving for the whole surface (as in manifold learning for instance). This latter property is very useful in designing manifold learning algorithms as well as dimension reduction techniques. The definition also easily handles bifurcations, graphs, trees that underlie the structure; most manifold learning algorithms assume a globally smooth manifold and fail to handle singularities easily or rigorously.